As part of its ensemble module, sklearn provides an AdaBoostClassifier implementation that supports two or more classes. The code examples for this section are in the notebook gbm_baseline that compares the performance of various algorithms with a dummy classifier that always predicts the most frequent class.
We need to first define a base_estimator as a template for all ensemble members and then configure the ensemble itself. We'll use the default DecisionTreeClassifier with max_depth=1—that is, a stump with a single split. The complexity of the base_estimator is a key tuning parameter because it depends on the nature of the data. As demonstrated in the previous chapter, changes to max_depth should be combined with appropriate regularization constraints using adjustments to, for example, min_samples_split, as shown in the following code:
base_estimator = DecisionTreeClassifier(criterion='gini',
splitter='best',
max_depth=1,
min_samples_split=2,
min_samples_leaf=20,
min_weight_fraction_leaf=0.0,
max_features=None,
random_state=None,
max_leaf_nodes=None,
min_impurity_decrease=0.0,
min_impurity_split=None)
In the second step, we'll design the ensemble. The n_estimators parameter controls the number of weak learners and the learning_rate determines the contribution of each weak learner, as shown in the following code. By default, weak learners are decision tree stumps:
ada_clf = AdaBoostClassifier(base_estimator=base_estimator,
n_estimators=200,
learning_rate=1.0,
algorithm='SAMME.R',
random_state=42)
The main tuning parameters that are responsible for good results are n_estimators and the base estimator complexity because the depth of the tree controls the extent of the interaction among the features.
We will cross-validate the AdaBoost ensemble using a custom 12-fold rolling time-series split to predict 1 month ahead for the last 12 months in the sample, using all available prior data for training, as shown in the following code:
cv = OneStepTimeSeriesSplit(n_splits=12, test_period_length=1, shuffle=True)
def run_cv(clf, X=X_dummies, y=y, metrics=metrics, cv=cv, fit_params=None):
return cross_validate(estimator=clf,
X=X,
y=y,
scoring=list(metrics.keys()),
cv=cv,
return_train_score=True,
n_jobs=-1, # use all cores
verbose=1,
fit_params=fit_params)
The result shows a weighted test accuracy of 0.62, a test AUC of 0.6665, and a negative log loss of -0.6923, as well as a test F1 score of 0.5876, as shown in the following screenshot:
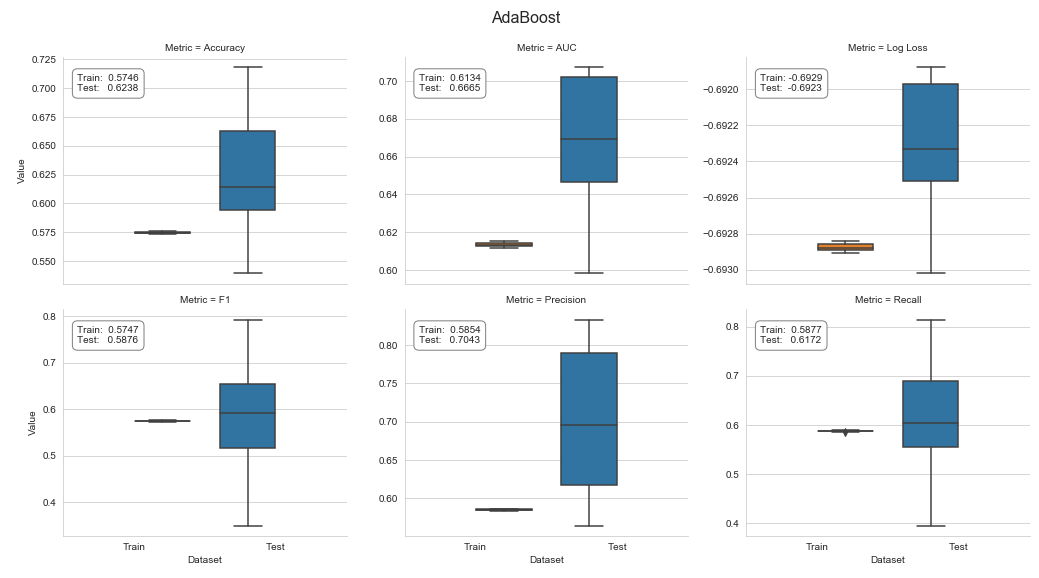
See the companion notebook for additional details on the code to cross-validate and process the results.
